- RAMOT at Tel Aviv University Ltd.
- From Israel
- Responsive
- Innovative Products and Technologies
Summary of the technology
Synthetic lethality occurs when a perturbation of two nonessential genes is lethal. This phenomenon offers a unique opportunity to develop selective anticancer drugs that will target a gene whose Synthetic Lethal (SL)-partner is inactive only in the cancer cells. Towards the realization of this potential, screening technologies have been developed to detect SL-interactions in model organisms and in human cell lines. However, currently their scope falls short compared to the large spectrum of interactions that need to be surveyed in cancer. Here we present a data-driven computational pipeline for the genome-wide identification of SL-interactions in cancer, by analyzing large volumes of genomic profiles of cancer cell lines and clinical samples. Our approach is first shown to successfully capture the SL-partners of tumor suppressors and oncogenes, according to six published experimental SL-screens. Second, we construct the first genome-wide network of SL-interactions in cancer and demonstrate its utility in predicting gene essentiality in a cancer-specific manner. Third, we show that, as predicted, underexpression of SL-partners has a strong positive prognostic value for cancer survival. Fourth, we identify synthetic lethality arising from gene over-activation and show that it can be utilized to predict anticancer drug efficacy. Lastly, we develop a complementary computational approach that by harnessing the obtained SL-networks identifies the most suitable treatment per patient, and identifies novel anticancer drug-targets, providing drug repurposing leads. Our approach provides a solid basis guiding future experimental studies aimed at developing new, selective anticancer therapies.
Project ID : 2-2013-713
Details of the Technology Offer
Technology
A universal computational framework for identifying molecular pathways underlying cancer viability and resistance. Our approach is applied to mining large collections of cancer patients’ data and identifying classes of genetic interactions termed synthetic lethality (SL).

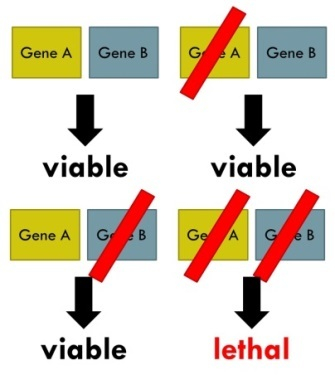
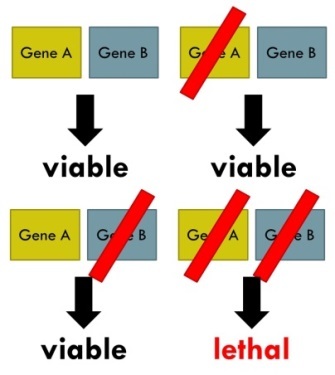
Synthetic lethality - The diagram above shows how a combination of mutations in two or more genes leads to cell death (lethality), whereas a mutation in only one of these genes is said to be viable.
The Need: A personalized, customized treatment targeting each tumor's weak spot
Each cancer tumor has its own genetic profile and therefore reacts differently to the treatments used to eliminate it. The world of personalized medicine needs technologies that can link a specific tumor with a specific treatment.
Often, tumors develop resistance to the drugs used to treat them. There is a great unmet need in predicting which tumors have a high probability of developing such resistance, as well as ways of preventing such resistance from occurring in the first place.
Technology today has reached the point where data generation is not a problem. The challenge is how to use sophisticated algorithms to analyze big data and identify the precise genes to be targeted in order to eliminate specific subsets of tumors.

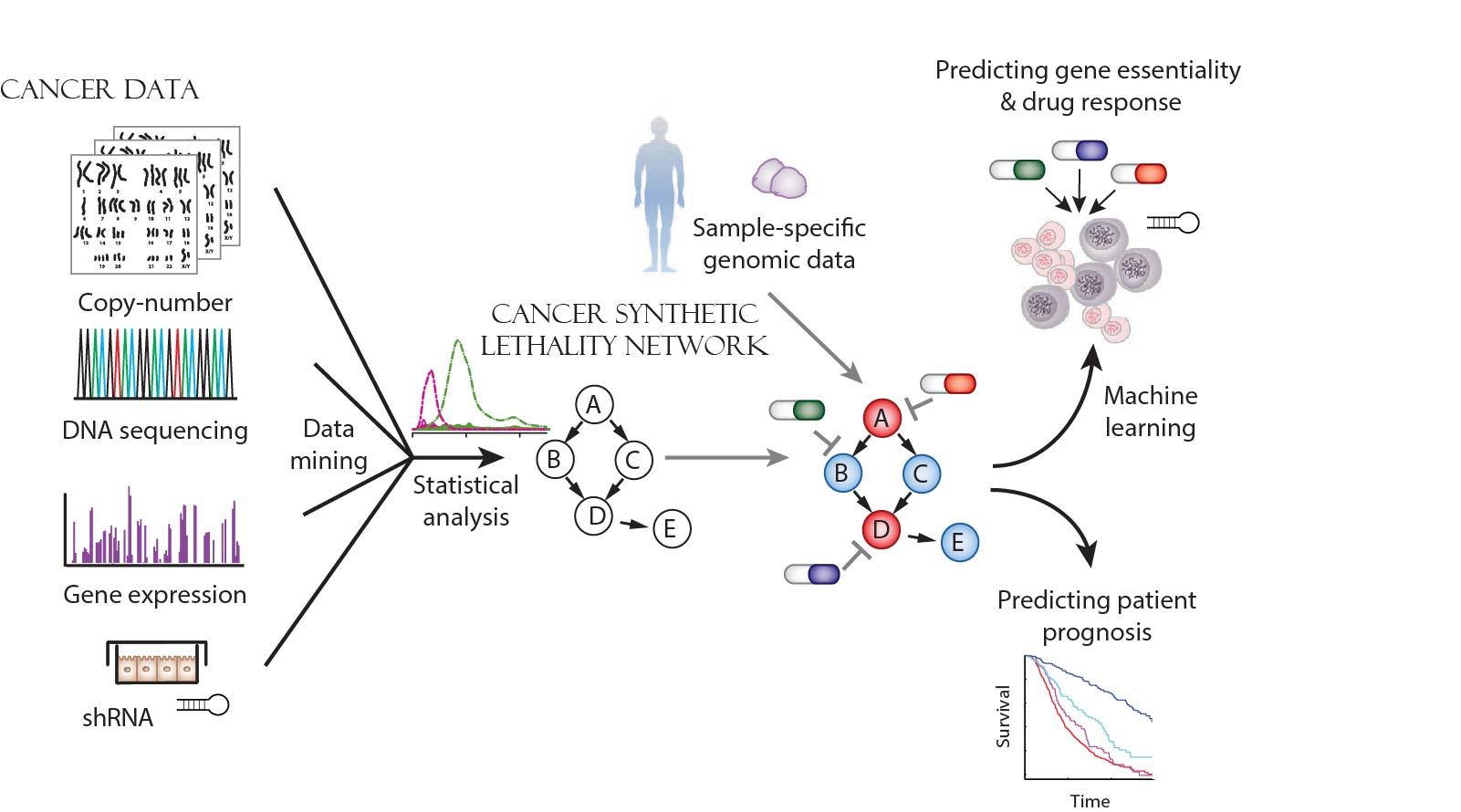
How it works – statistical analysis is performed on various data from actual tumors and a network of gene interactions is formed. Then, genetic data from a specific patient's tumor is fed into the network and the tumor's weak spot is identified for different applications
Applications
A 'personalized medicine' data analysis platform for predicting tumor weak spots for the following applications:
Stratification of clinical trial patients – Clinical trials can be designed to include only patients whose tumor is predicted to interact with the proposed drug.
Prediction of drug response - A diagnostic tool to assist oncologists in choosing the correct course of treatment. The technology predicts positive/negative drug response based on the analysis of the patient’s tumor.
Drug repurposing – Using this technology, FDA approved drugs (not necessarily cancer drugs) with a known 'Mode of Action' can be repurposed to treat subsets of certain cancers.
New targets for drug development – Using this technology, a large number of new targets for drug development can be identified.
Novel adjuvants that can prevent the tumor from developing drug resistance – Based on the predicted type of resistance, this technology can match a relevant adjuvant that will block the tumor from developing the specific resistance.
Stage of Development
The algorithms were validated using large-scale prediction models and actual treatment of cancer cells.
Patents
US patent application filed - CANCER PROGNOSIS AND THERAPY BASED ON SYNTHETIC LETHALITY
Provisional patent filed in December 2015 - SLICK: A clinically based pipeline for the identification and utilization of synthetic lethal interactions to advance cancer therapy
Supporting Publications
Predicting cancer-specific vulnerability via data-driven detection of synthetic lethality. Cell. 2014 Aug 28; 158(5):1199-209.