Showing 1 to 15 of 2075 results


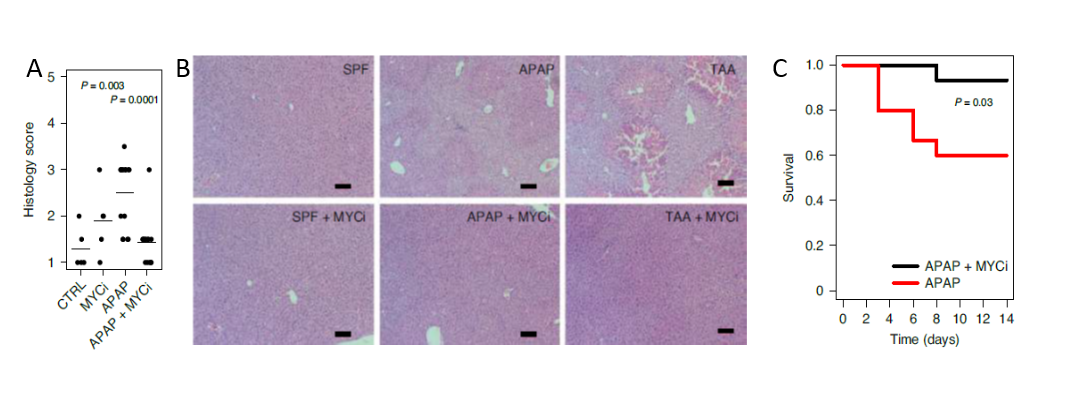
ProteinGAN – deep learning framework for rapid generation of highly-functional synthetic enzyme libraries
Knowhow and Research output Vilnius University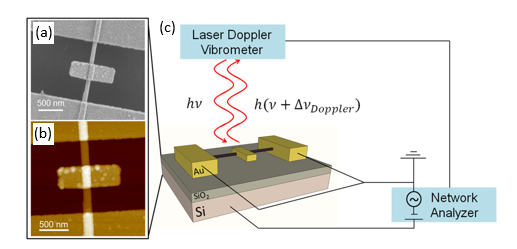

Inertial NEMS for Nanosensor Applications Based on Inorganic Nanotubes
Patents for licensing Yeda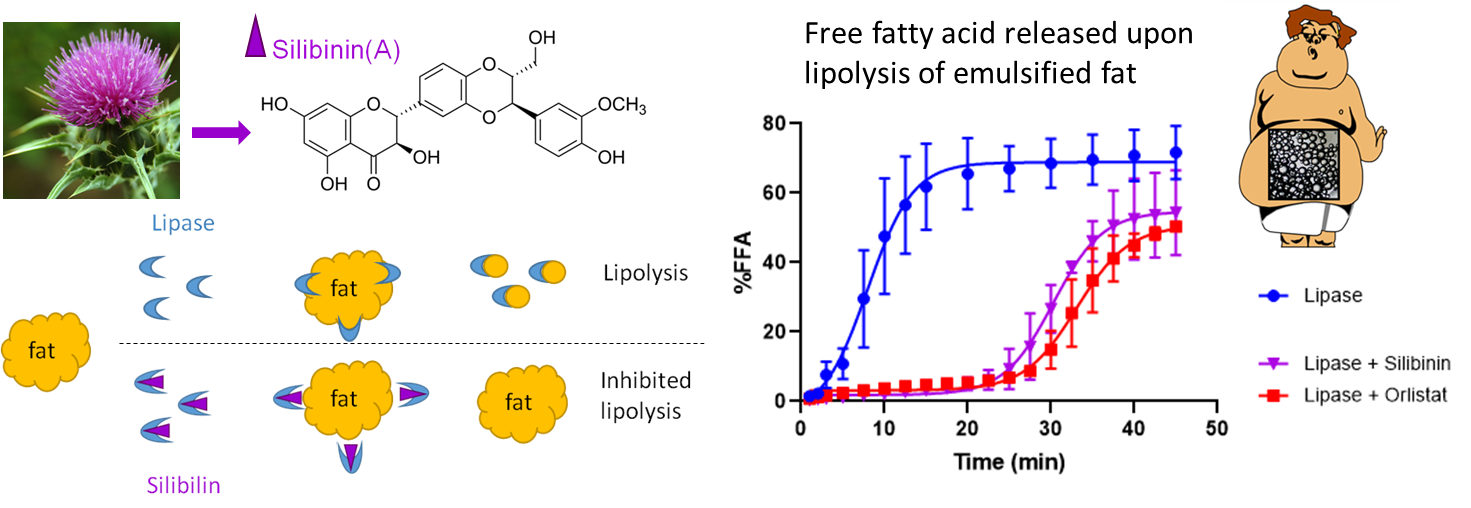

Compound for the treatment of obesity and hyperlipidemia
Patents for licensing Universidad de Granada

Iron Catalyzed Ring-Opening Metathesis Polymerization (ROMP)
Patents for licensing Yeda

Pharmaceutical composition for the treatment of cystic fibrosis
Patents for licensing UNIVERSIDAD DE BURGOS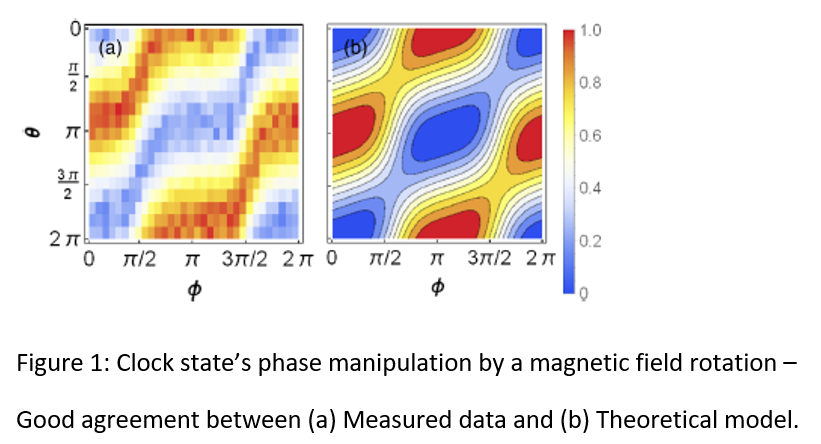

Atomic Magnetometer with Increased Coherence Time for Improved Magnetic Fields
Patents for licensing Yeda

Artificial reefs for safe and sustainable marine habitat regeneration
Patents for licensing Universidad de Alicante

New Antiviral Drugs from Bacterial Natural Products
Patents for licensing Yeda

Super Porous Adsorbent of Heavy metals for Water Purification
Patents for licensing ICN2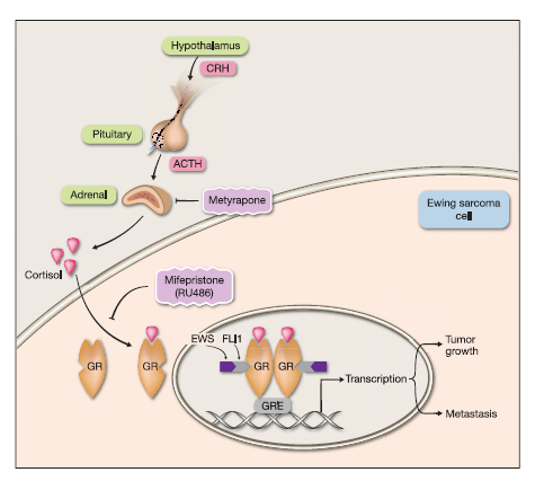

Treatment for Ewing Sarcoma Using ETS Proteins
Patents for licensing Yeda![Automatic Tool for Scouring the Internet and Generating Names Linked to Illegal Activities to Facilitate Detection and Reporting o[…]](https://static3.innoget.com/images/bgpremium.png)

Automatic Tool for Scouring the Internet and Generating Names Linked to Illegal Activities to Facilitate Detection and Reporting o[…]
Innovative Products and Technologies Northeastern University's Center for Research Innovation

Biomelanin synthesised from Fungi
Innovative Products and Technologies QuickGun LifeScience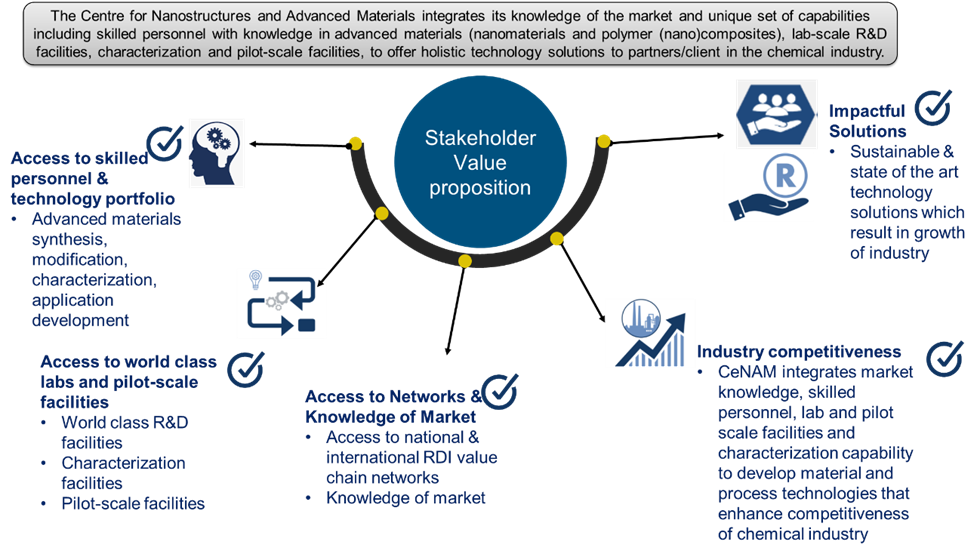

Access to worldclass laboratory, pilot-scale facilities, skilled personnel and technology portfolio.
Research Services and Capabilities CSIR

Anti importin beta1 monoclonal antibody
Patents for licensing Yeda