Showing 1 to 15 of 2075 results


Efficient and selective magnetic adsorbent for removal of purine compounds and nucleic acids from liquid matrices
Patents for licensing UATEC - Unidade de Transferência de Tecnologia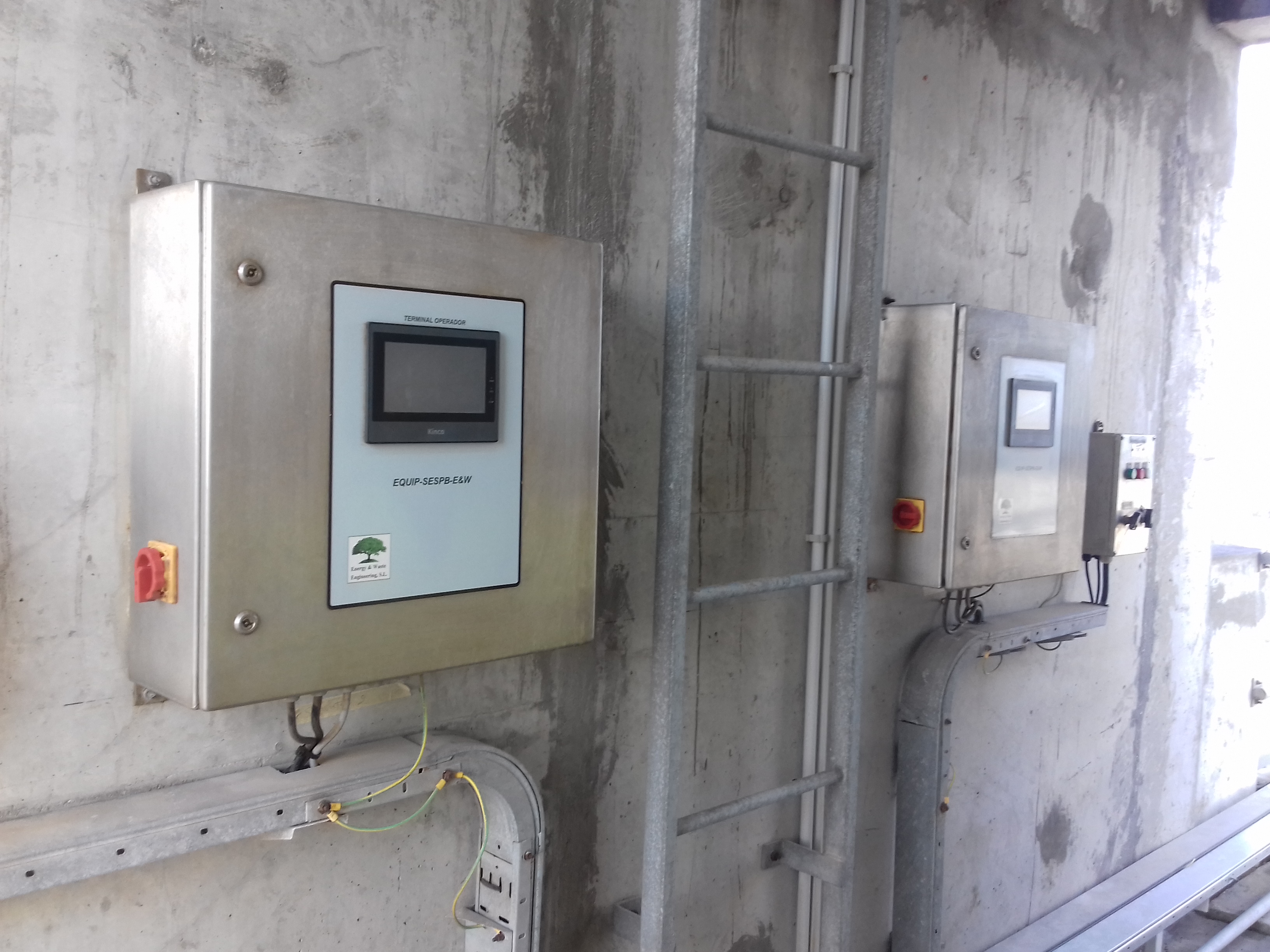

BTS-Foam and Sediments Removal
Innovative Products and Technologies Biogas & Gases Technologies

A Software System for the Microbial Source Tracking Problem
Patents for licensing Universitat Politècnica de Catalunya - UPC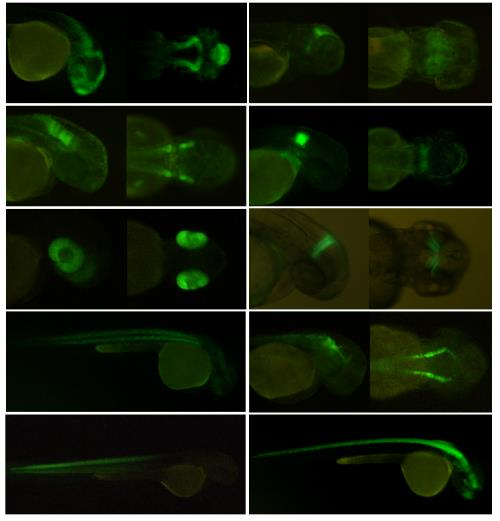

Kit for the systematic generation of transgenic lines for transcriptomic analyses
Patents for licensing Consejo Superior de Investigaciones Científicas

Bioresorbable polymer composite for bone-soft tissue fixation.
Innovative Products and Technologies Centre For Future (CFF)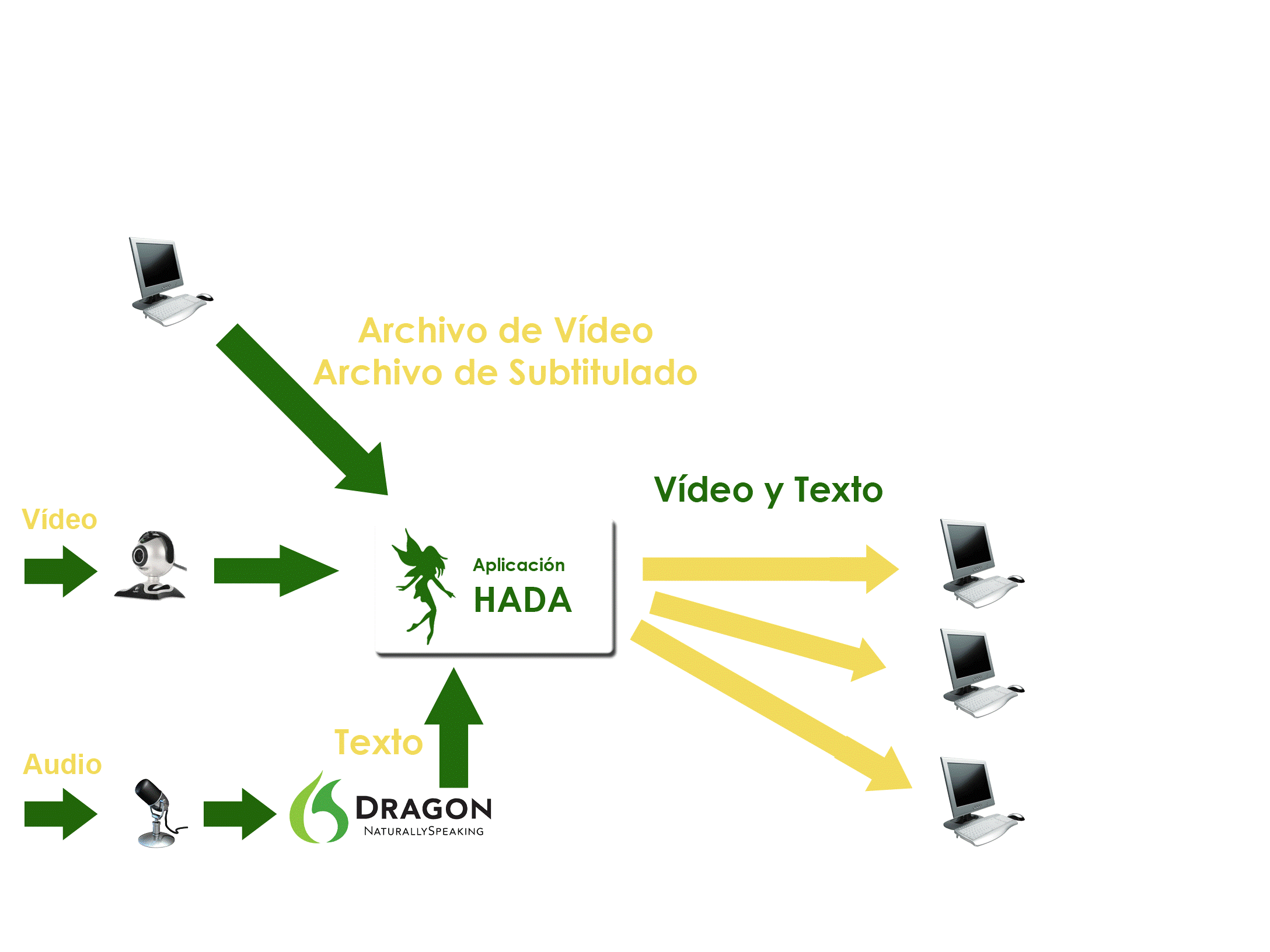

HADA (Hearing Impaired Assistance Tool). Real-time text and video streaming for the hearing impaired people
Patents for licensing UNIVERSIDAD DE BURGOS

Direct sensor for electron microscopy with in-pixel energy measurement
Patents for licensing Consejo Superior de Investigaciones Científicas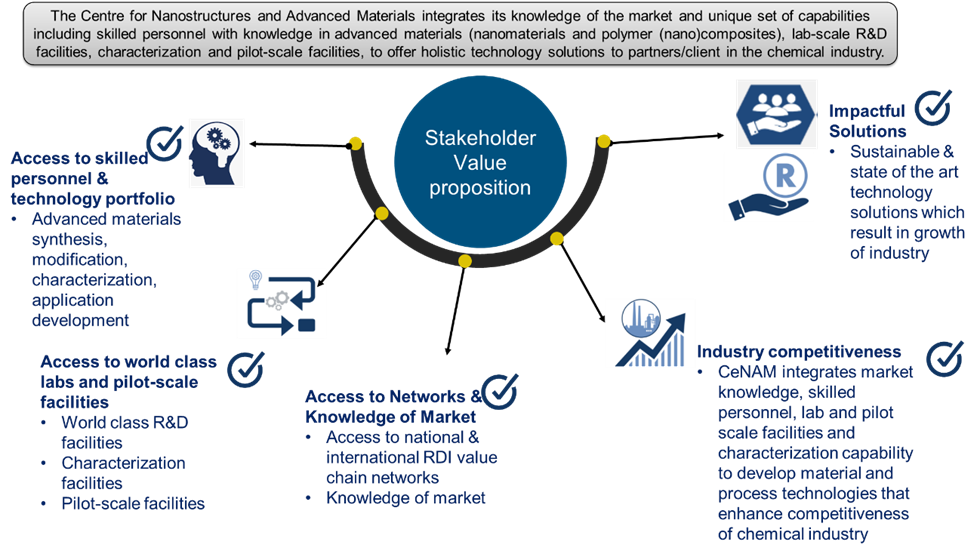

Access to worldclass laboratory, pilot-scale facilities, skilled personnel and technology portfolio.
Research Services and Capabilities CSIR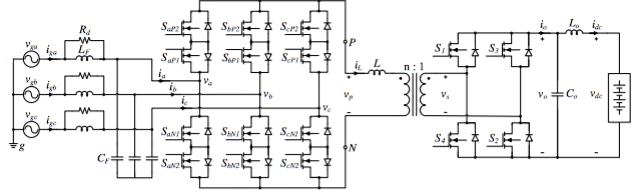

a single-stage, bidirectional and high-frequency isolated power conversion system
Patents for licensing INESC TEC

Dual/Triple Sump pump controller with optional bluetooth or Wifi connectivity
Innovative Products and Technologies Chicagoland Automation

High resolution paterning in organic semiconductors
Patents for licensing Consejo Superior de Investigaciones Científicas

Nanofaber
Investment Opportunities in Startups and Spinoffs CINBIO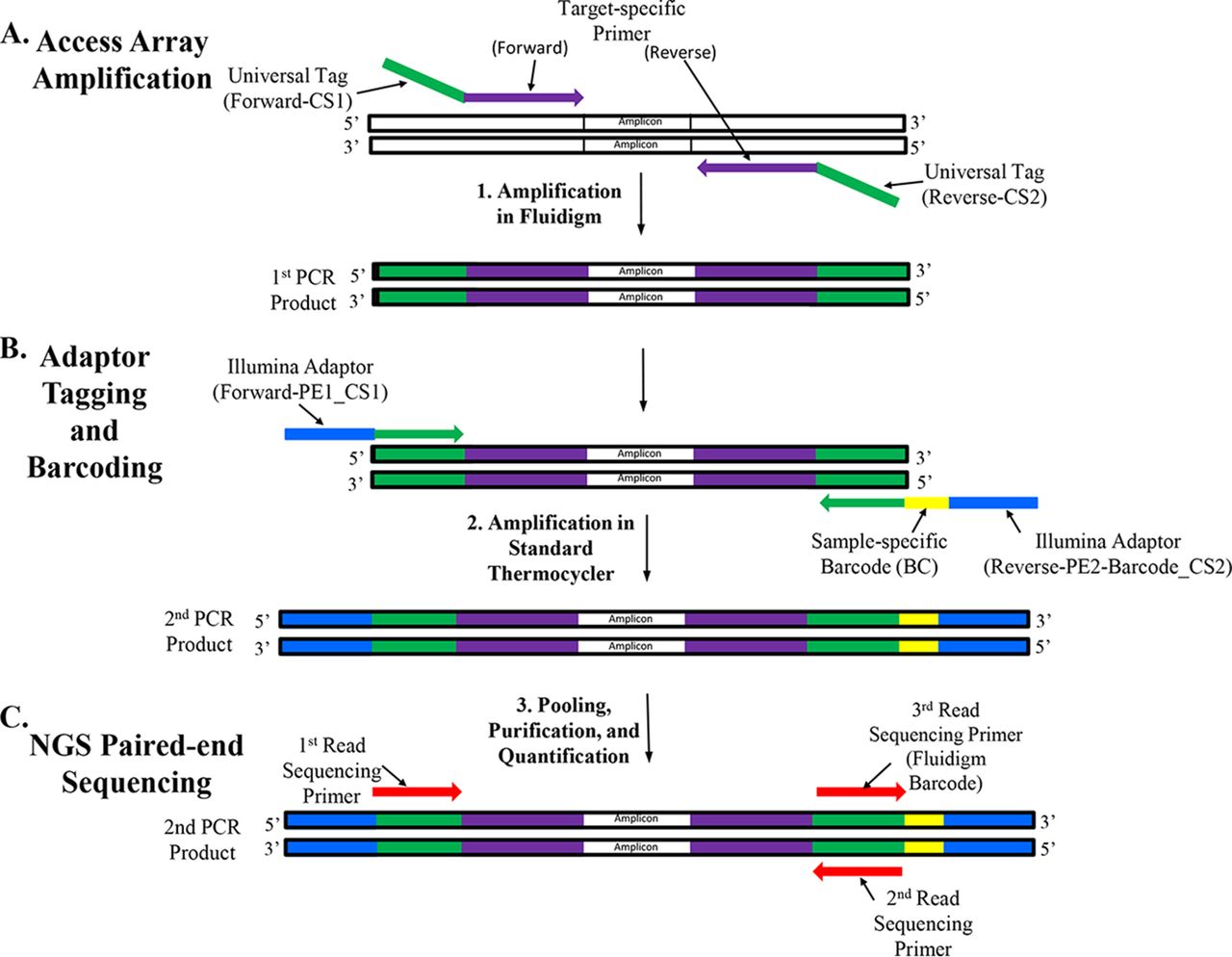

Amplicon Sequencing-Microbialtec Research
Research Services and Capabilities Microbialtec Division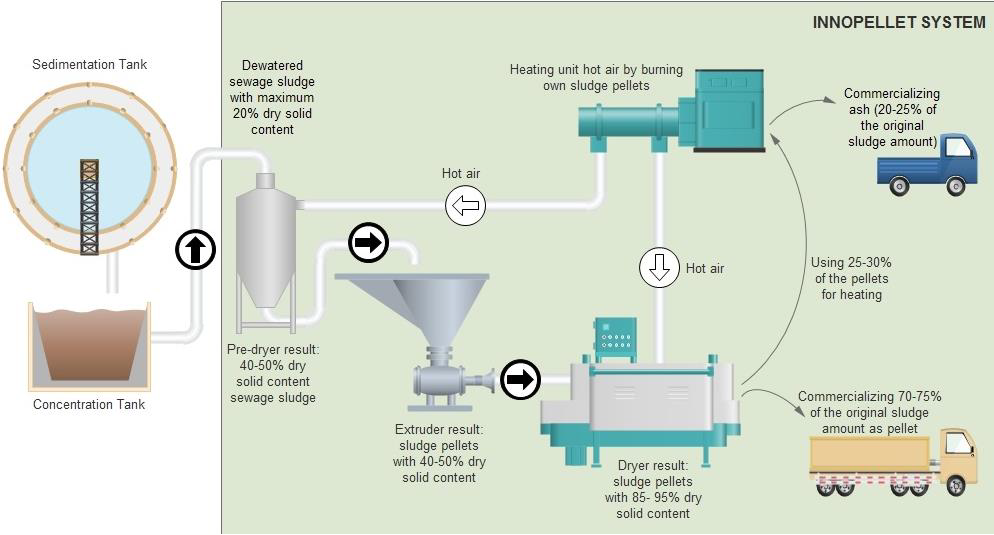

An Innovative Technology to Effectively Utilize and Recycle Sewage Sludge
Innovative Products and Technologies Laser Consult Ltd.![New methods for the detection and discrimination of contaminants and organic metabolites of high environmental impact by means of […]](https://static9.innoget.com/uploads//79d0a575a1a9ecbca8a1f563aaae257a93d651a5.png)

