Showing 1 to 15 of 2075 results


CHEMIOTHERAPY: IRINOTECAN DRUG MONITORING
Patents for licensing National Cancer Institute CRO Aviano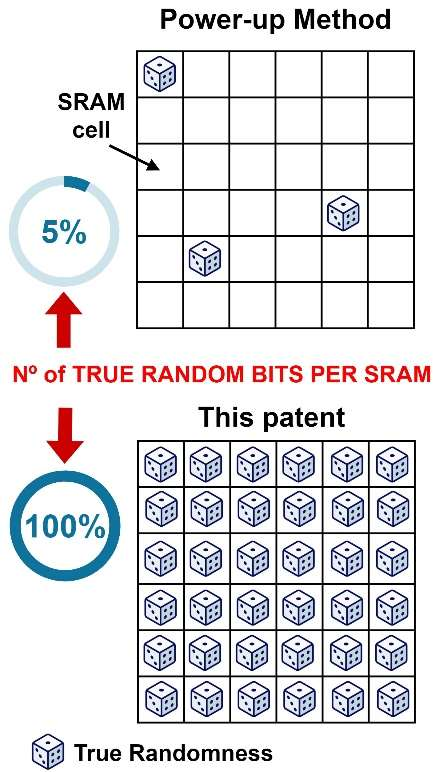

Method and device for generating true random numbers
Patents for licensing CSIC - Consejo Superior de Investigaciones Científicas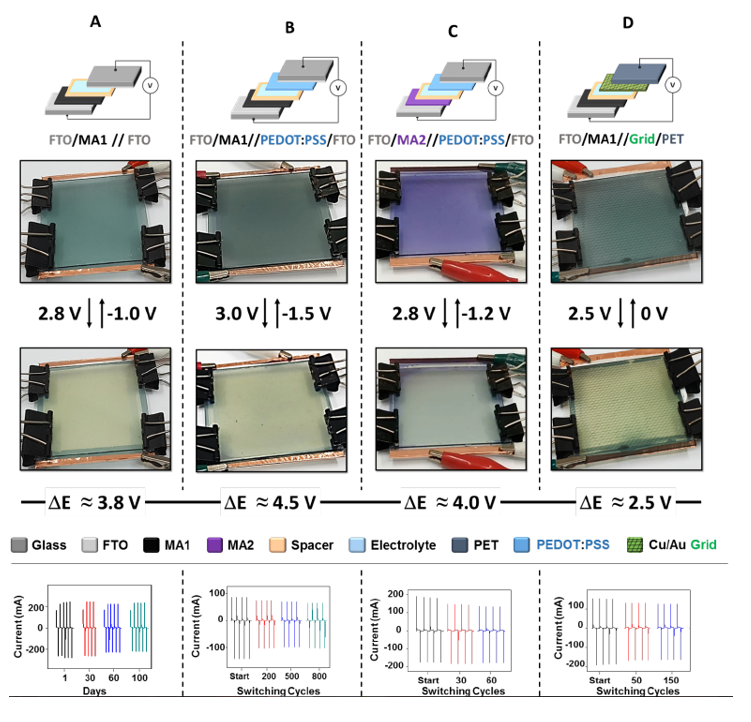

High-Performance Electrochromic Devices
Patents for licensing Yeda
Direct Solar Lighting Technology
Investment Opportunities in Startups and Spinoffs i-EL Technologies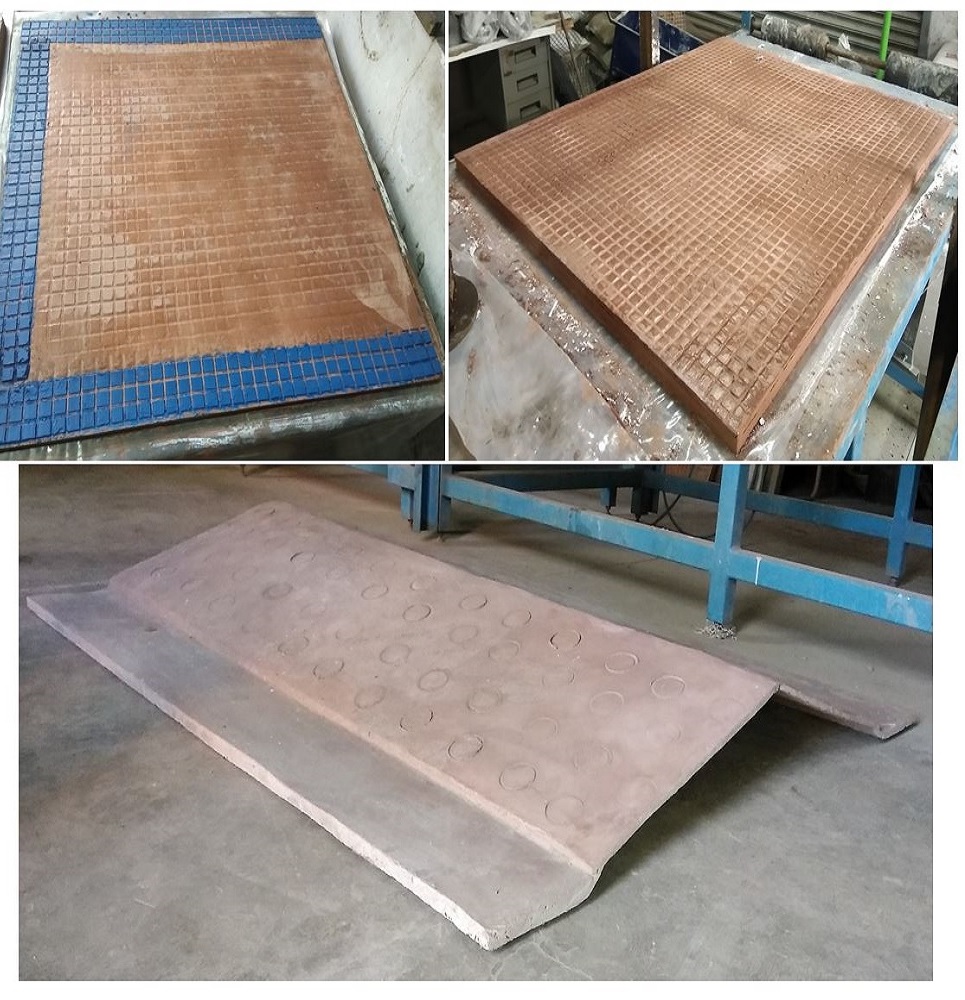

Textile Reinforced Concrete Prototyping Technology
Knowhow and Research output Steinbeis Centre for Technology Transfer India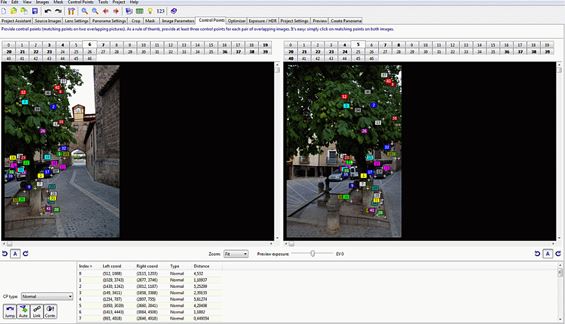

Design and development of virtual and/or interactive guides/tours
Research Services and Capabilities UNIVERSIDAD DE BURGOS

Prefabricated of cement with fire resistant polymeric residues
Patents for licensing UNIVERSIDAD DE BURGOS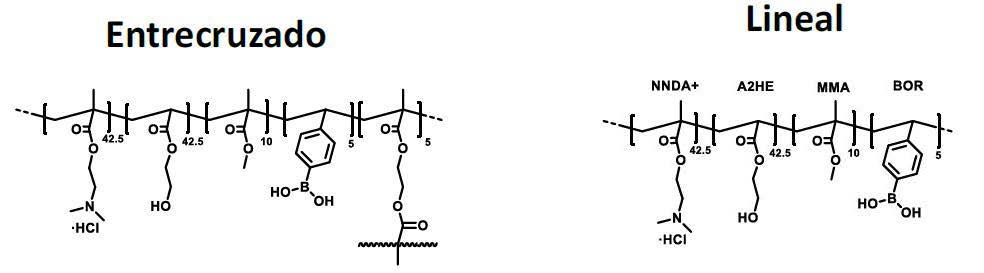

Polymer capable of extracting aromatic alcohols from honey
Patents for licensing UNIVERSIDAD DE BURGOS

Graphene-based electroactive fluids for Energy Storage
Patents for licensing Consejo Superior de Investigaciones Científicas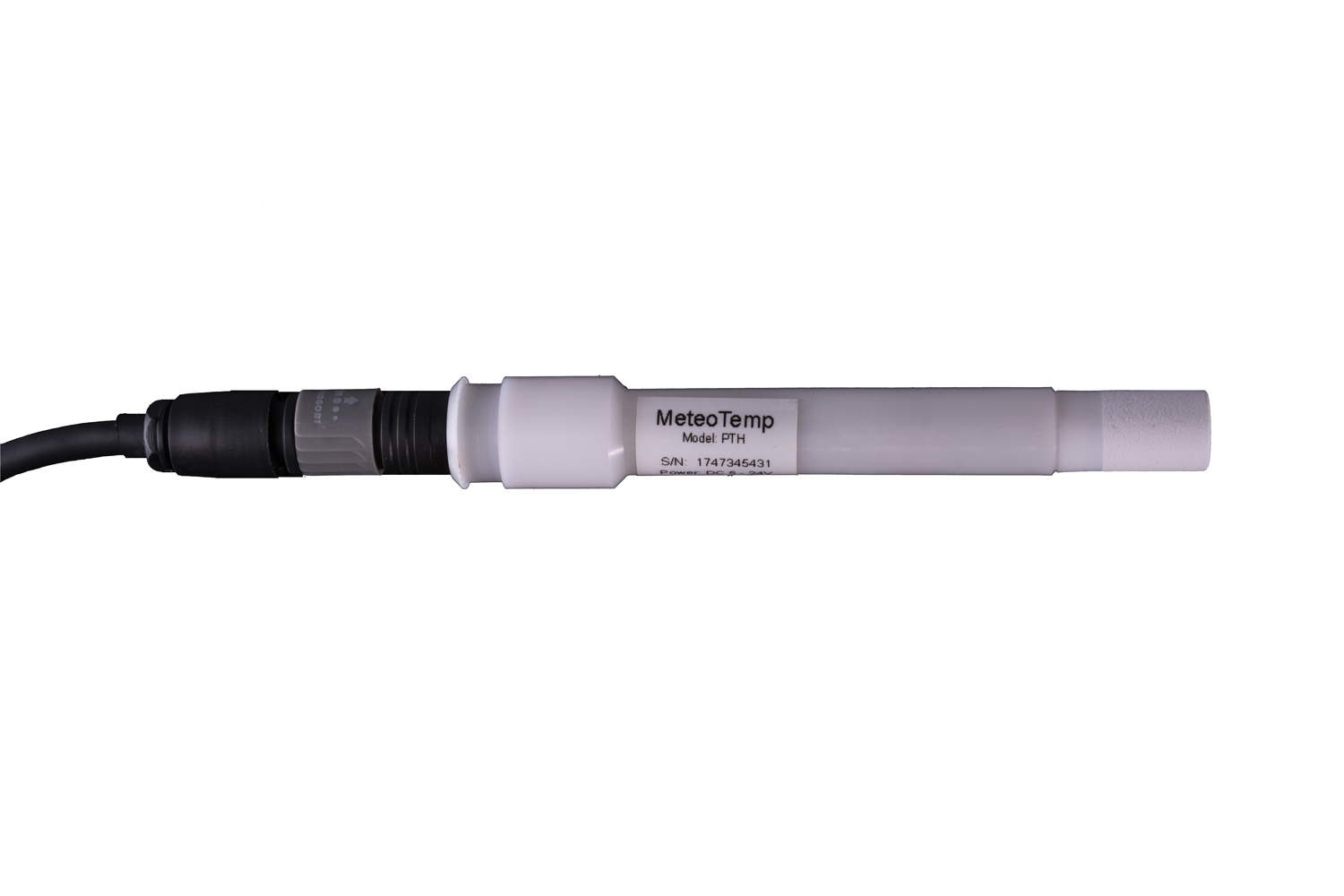

Ultra low power MODBUS Temperature, Humidity, Pressure sensor
Innovative Products and Technologies Barani Design

Vanillin (a Greener and cost-effective conversion of lignin and ferulic acid into vanillin)
Patents for licensing University of Ljubljana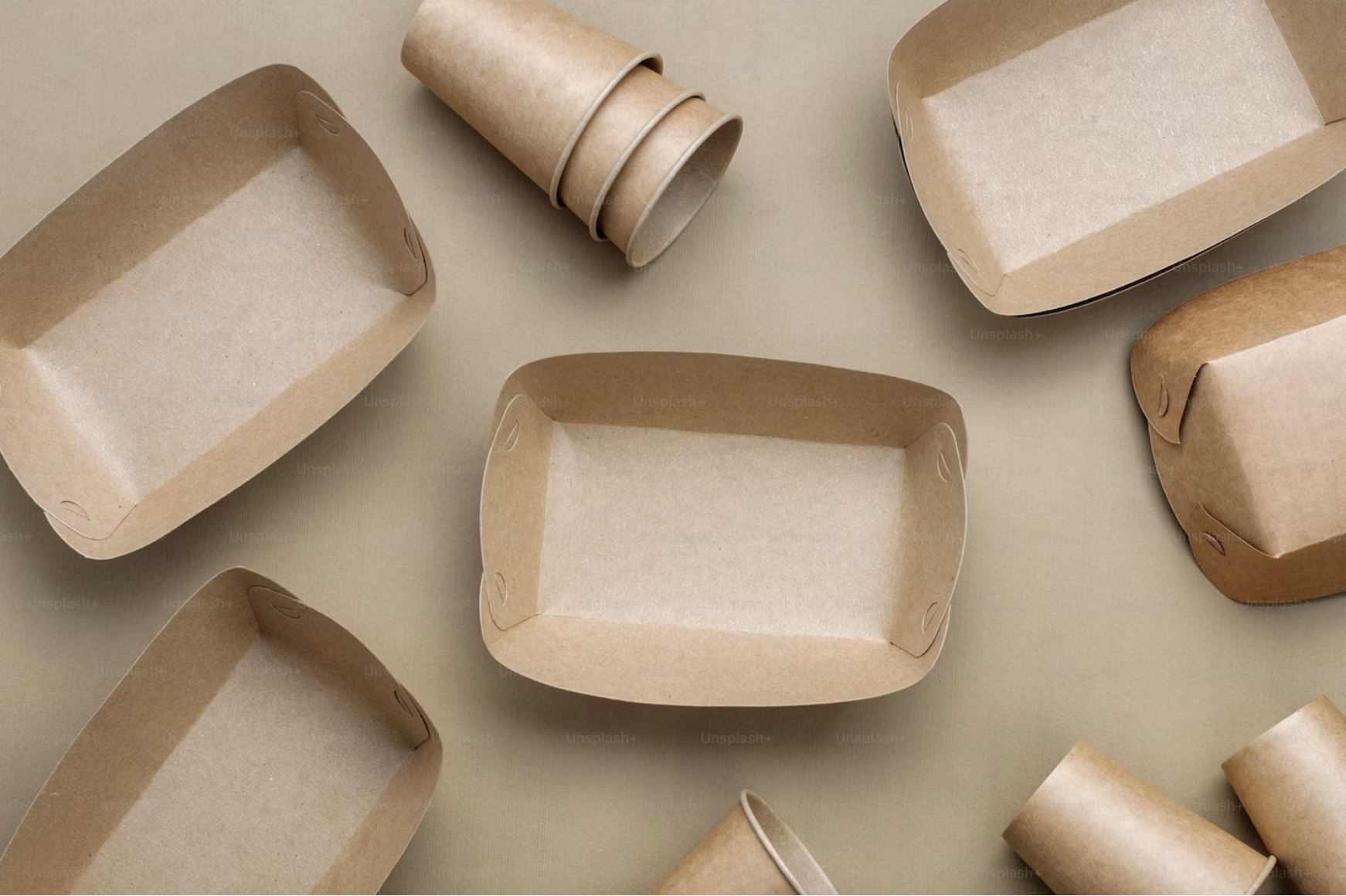

Reinforced packaging paper
Patents for licensing CSIC - Consejo Superior de Investigaciones Científicas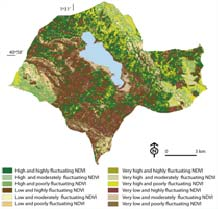

Environmental indices through LandSAT images
Patents for licensing Consejo Superior de Investigaciones Científicas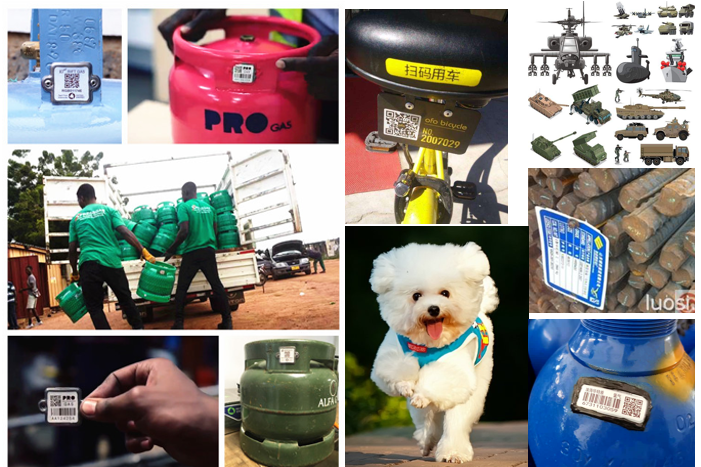

Gas cylinders to have QR codes to stop theft
Innovative Products and Technologies XiangKang

